Everything should be made as simple as possible, but not simpler.
— Albert Einstein
Original 3D Data
Principal Component Analysis is a dimensionality reduction technique commonly used in machine learning. In simple words, it rewrites data in a new coordinate system so that the most important patterns come first and the noise comes last.
Linear Algebra Prerequisites
Let the dataset contain samples and features. We write it as a matrix , where each row is one sample.
PCA looks for a new orthonormal basis for this feature space. A basis is just a set of directions that can build every vector in the space. Orthonormal means two things: the directions are perpendicular, and each one has length .
Tiny analogy: if the original axes are a fixed grid on the floor, PCA is like rotating that grid until it lines up with the longest stretch of the crowd. The grid itself changes, but the crowd does not.
If contains the principal directions, then projecting the centered data onto those directions gives a new representation:
where is the centered data matrix.
This is a change of basis. The original axes may be convenient for storage, but they are not always the best axes for describing variation. PCA replaces them with axes aligned to the data itself.
That makes the transformed coordinates easy to interpret and mathematically clean.

The Problem
Imagine you are building a fraud detector for credit card transactions. Your dataset contains many features: merchant category, country, time of day, amount, device type, transaction channel, and so on. After exploratory analysis, you notice that several features move together. For example, online transactions may be more likely to be fraudulent, and unusually large amounts may be suspicious when compared with a user’s normal spending pattern.
This creates two problems. First, the data has redundancy: multiple features may be telling you almost the same story. Second, the data may have noise: small fluctuations that do not help prediction. If the feature count is tiny, you can often inspect things manually. In real-world problems, though, the number of features can be large enough that this becomes impractical. PCA helps by compressing the data while keeping as much useful variation as possible.
This is where PCA comes in. You choose how many components to keep, and PCA finds the directions in which the data varies the most. Those directions are ordered from most informative to least informative, and the data is projected onto the top few. The result is a smaller representation that usually retains the structure of the original dataset surprisingly well.
The key idea is geometric: instead of asking “which features are most useful?”, PCA asks “along which directions does this cloud of points spread out the most?”
Variance and Covariance
To find the best basis, we need a way to measure spread. Variance tells us how much one feature changes by itself. Covariance tells us how two features change together.
For centered data , the covariance matrix is
Each entry measures how feature and feature vary together.
If we can find a transformation that makes the covariance matrix diagonal, then the new features will be uncorrelated. That means each new axis captures its own piece of variation, without duplicating what another axis already explains.

The Proof
Start with the covariance matrix of the centered data:
Because is symmetric, the spectral theorem guarantees an orthogonal eigendecomposition:
where:
- contains the eigenvectors of
- is a diagonal matrix of eigenvalues
There is also an optimization view of the same idea. Suppose we choose a unit vector and project every centered sample onto it. The variance of those projected points is
with the constraint
Maximizing under that constraint gives the first principal component. Using a Lagrange multiplier, the stationary points satisfy
so the best direction is an eigenvector of the covariance matrix. The largest eigenvalue gives the direction with the greatest variance.
The eigenvectors point in the directions of principal variation, and the eigenvalues tell us how much variance lies in each direction.
Now project the centered data onto those eigenvectors:
If we keep all components, the covariance of the projected data is
This is diagonal. So the transformed coordinates are uncorrelated, and each axis captures a distinct amount of variance.
If we keep only the first eigenvectors, we get a lower-dimensional projection that preserves the most variance available in directions. That is the core of PCA.
2D PCA Projection
Implementation
Now that the geometry is clear, the implementation becomes straightforward.
First, center the data by subtracting the mean of each feature. PCA cares about variation around the data’s own center, not around the origin. Without centering, the first component can be dominated by an arbitrary offset instead of true structure.
Think of a crowd in a field. If you ask for the direction in which the crowd is most spread out, you would naturally measure that around the center of the crowd. If you instead measured everything from a random flag far away, you would distort the result.
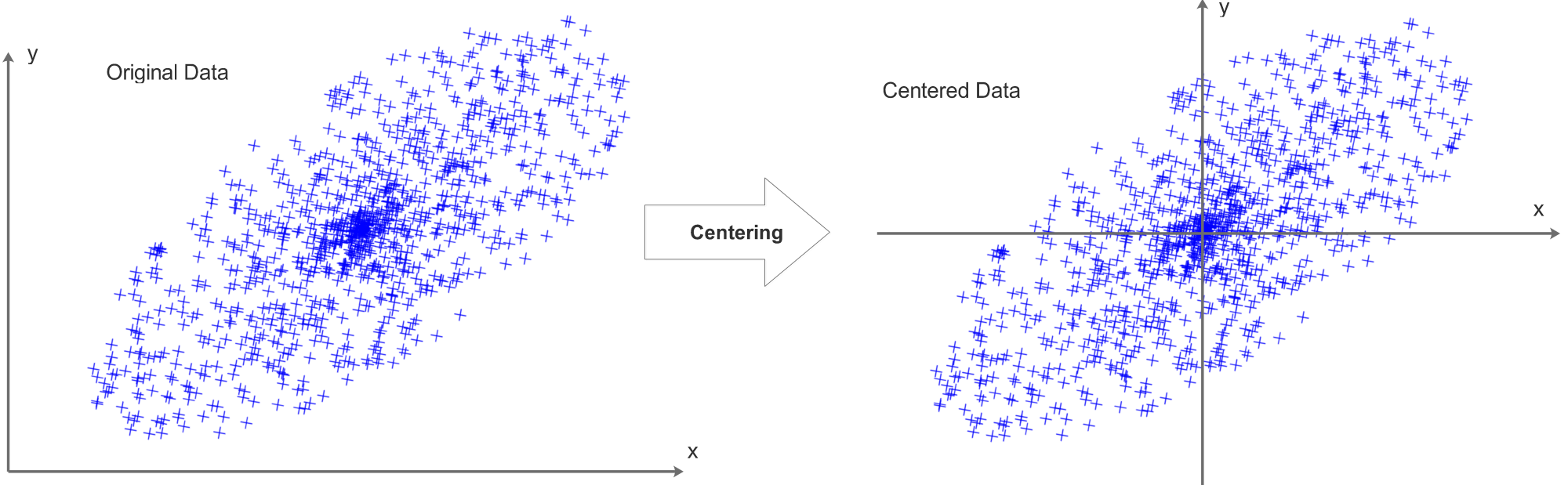
After centering, we compute the covariance matrix and perform eigendecomposition. Then we sort the eigenpairs, keep the top directions, and project the data.
Here is a complete, minimal implementation you can run directly:
import numpy as np
class PCA:
def __init__(self, n_components: int):
if n_components <= 0:
raise ValueError("n_components must be positive")
self.n_components = n_components
# Learned during fit
self.mean_ = None
self.components_ = None
self.explained_variance_ = None
self.explained_variance_ratio_ = None
def fit(self, X: np.ndarray):
X = np.asarray(X, dtype=float)
if X.ndim != 2:
raise ValueError("X must be a 2D array of shape (n_samples, n_features)")
n_samples, n_features = X.shape
if n_samples < 2:
raise ValueError("Need at least 2 samples to compute covariance")
if self.n_components > n_features:
raise ValueError("n_components cannot exceed number of features")
# 1) Center data
self.mean_ = X.mean(axis=0)
X_centered = X - self.mean_
# 2) Covariance matrix
covariance = (X_centered.T @ X_centered) / (n_samples - 1)
# 3) Eigendecomposition for symmetric matrix
eigvals, eigvecs = np.linalg.eigh(covariance)
# 4) Sort by descending eigenvalue
order = np.argsort(eigvals)[::-1]
eigvals = eigvals[order]
eigvecs = eigvecs[:, order]
# 5) Keep top-k principal directions
# eigvecs columns are eigenvectors; components_ shape: (k, n_features)
self.components_ = eigvecs[:, : self.n_components].T
self.explained_variance_ = eigvals[: self.n_components]
total_var = eigvals.sum()
self.explained_variance_ratio_ = self.explained_variance_ / total_var
return self
def transform(self, X: np.ndarray) -> np.ndarray:
if self.mean_ is None or self.components_ is None:
raise RuntimeError("Call fit before transform")
X = np.asarray(X, dtype=float)
X_centered = X - self.mean_
# Project to lower-dimensional space: (n_samples, n_features) @ (n_features, k)
return X_centered @ self.components_.T
def fit_transform(self, X: np.ndarray) -> np.ndarray:
return self.fit(X).transform(X)
def inverse_transform(self, Z: np.ndarray) -> np.ndarray:
if self.mean_ is None or self.components_ is None:
raise RuntimeError("Call fit before inverse_transform")
Z = np.asarray(Z, dtype=float)
# Reconstruct approximation in original feature space
return Z @ self.components_ + self.mean_Example usage:
rng = np.random.default_rng(7)
X = rng.normal(size=(200, 5))
pca = PCA(n_components=2)
Z = pca.fit_transform(X)
print("Reduced shape:", Z.shape)
print("Explained variance:", pca.explained_variance_)
print("Explained variance ratio:", pca.explained_variance_ratio_)
print("Cumulative explained variance:", pca.explained_variance_ratio_.sum())This implementation gives the core PCA workflow end-to-end: fit, project, and reconstruct.
It is useful to track explained variance ratio and its cumulative sum. If the first few components explain most of the variance, PCA has compressed the data while preserving most of the structure.
An alternate implementation of PCA based on SVD in python can be found here.
Conceptual framing inspired by: A Tutorial on Principal Component Analysis (Shlens 2014).